A pre-trained model for volumetric (3D) segmentation of the spleen from CT image
This model is trained using the runner-up [1] awarded pipeline of the “Medical Segmentation Decathlon Challenge 2018” using the UNet architecture [2] with 32 training images and 9 validation images.
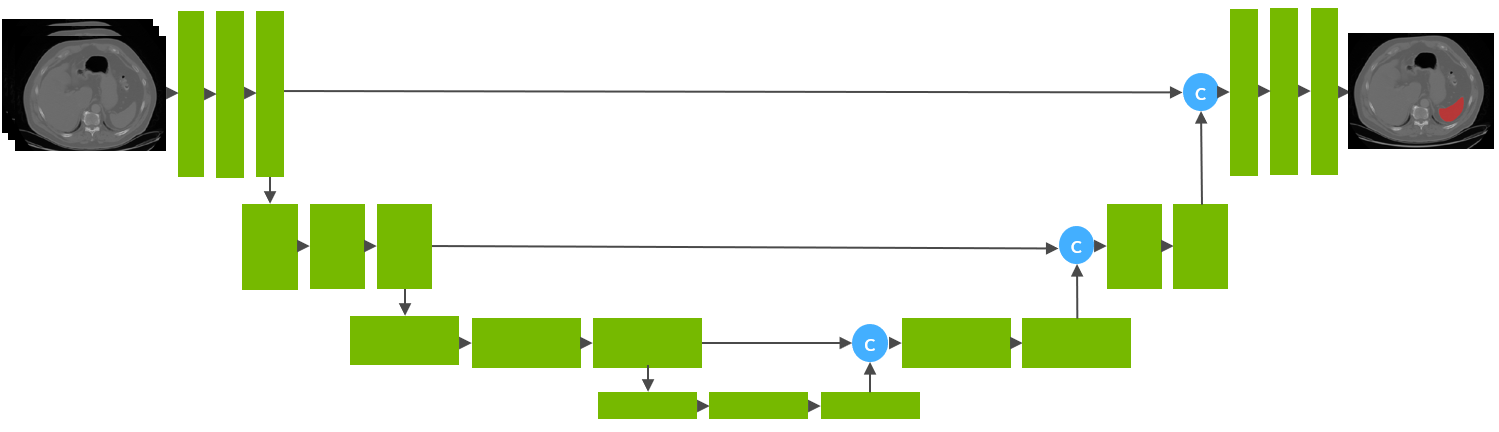
Data
The training dataset is the Spleen Task from the Medical Segmentation Decathalon. Users can find more details on the datasets at http://medicaldecathlon.com/ .
- Target: Spleen
- Modality: CT
- Size: 61 3D volumes (41 Training + 20 Testing)
- Source: Memorial Sloan Kettering Cancer Center
- Challenge: Large-ranging foreground size
Training configuration
The segmentation of spleen region is formulated as the voxel-wise binary classification. Each voxel is predicted as either foreground (spleen) or background. And the model is optimized with gradient descent method minimizing Dice + cross entropy loss between the predicted mask and ground truth segmentation.
The training was performed with the following:
- GPU: at least 12GB of GPU memory
- Actual Model Input: 96 x 96 x 96
- AMP: True
- Optimizer: Novograd
- Learning Rate: 0.002
- Loss: DiceCELoss
- Dataset Manager: CacheDataset
Memory Consumption Warning
If you face memory issues with CacheDataset, you can either switch to a regular Dataset class or lower the caching rate cache_rate in the configurations within range [0, 1] to minimize the System RAM requirements.
Input
One channel - CT image
Output
Two channels - Label 1: spleen - Label 0: everything else
Performance
Dice score is used for evaluating the performance of the model. This model achieves a mean dice score of 0.961.
Training Loss
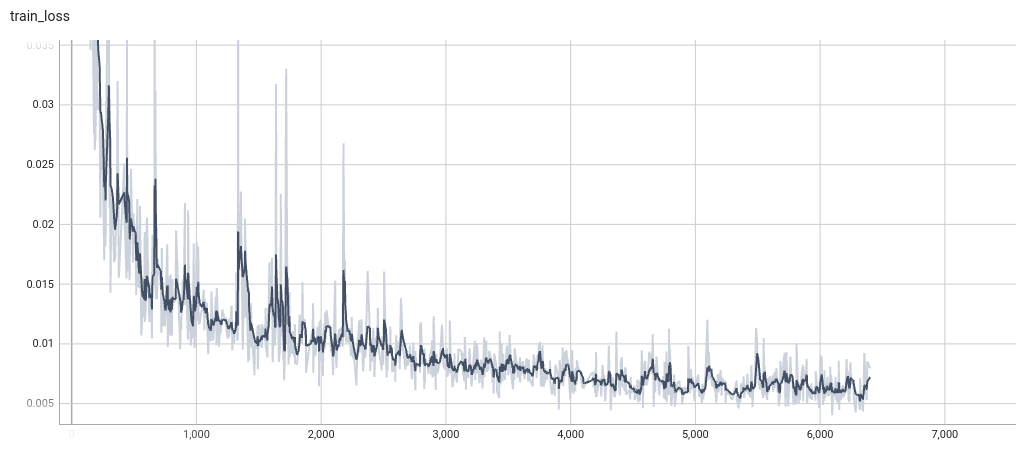
Validation Dice
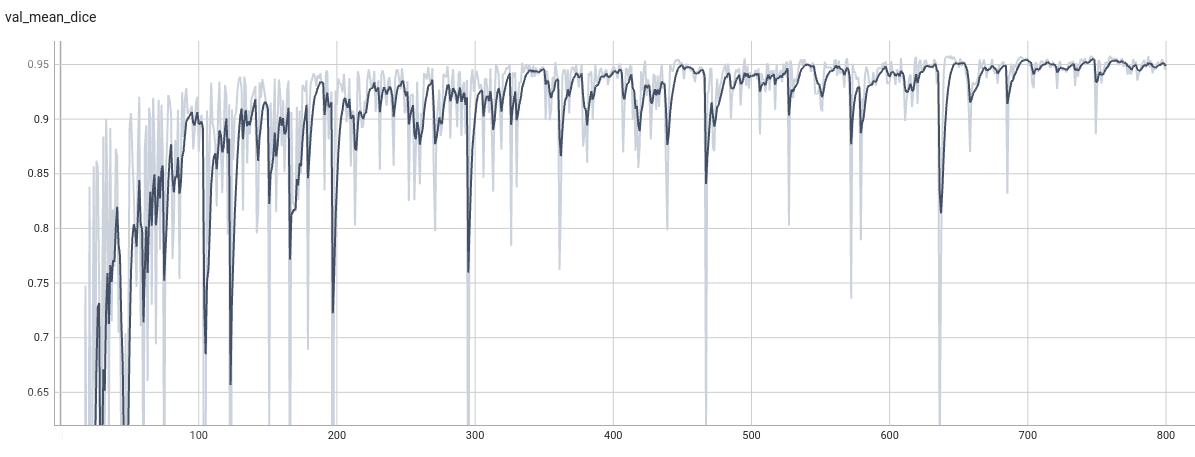
TensorRT speedup
The spleen_ct_segmentation bundle supports acceleration with TensorRT through the ONNX-TensorRT method. The table below displays the speedup ratios observed on an A100 80G GPU.
method
torch_fp32(ms)
torch_amp(ms)
trt_fp32(ms)
trt_fp16(ms)
speedup amp
speedup fp32
speedup fp16
amp vs fp16
model computation
6.46
4.48
2.52
1.96
1.44
2.56
3.30
2.29
end2end
1268.03
1152.40
1137.40
1114.25
1.10
1.11
1.14
1.03
Where: - model computation means the speedup ratio of model’s inference with a random input without preprocessing and postprocessing - end2end means run the bundle end-to-end with the TensorRT based model. - torch_fp32 and torch_amp are for the PyTorch models with or without amp mode. - trt_fp32 and trt_fp16 are for the TensorRT based models converted in corresponding precision. - speedup amp , speedup fp32 and speedup fp16 are the speedup ratios of corresponding models versus the PyTorch float32 model - amp vs fp16 is the speedup ratio between the PyTorch amp model and the TensorRT float16 based model.
Currently, the only available method to accelerate this model is through ONNX-TensorRT. However, the Torch-TensorRT method is under development and will be available in the near future.
This result is benchmarked under: - TensorRT: 8.5.3+cuda11.8 - Torch-TensorRT Version: 1.4.0 - CPU Architecture: x86-64 - OS: ubuntu 20.04 - Python version:3.8.10 - CUDA version: 12.1 - GPU models and configuration: A100 80G
MONAI Bundle Commands
In addition to the Pythonic APIs, a few command line interfaces (CLI) are provided to interact with the bundle. The CLI supports flexible use cases, such as overriding configs at runtime and predefining arguments in a file.
For more details usage instructions, visit the MONAI Bundle Configuration Page .
Execute training:
python -m monai.bundle run --config_file configs/train.json
Please note that if the default dataset path is not modified with the actual path in the bundle config files, you can also override it by using --dataset_dir :
python -m monai.bundle run --config_file configs/train.json --dataset_dir <actual dataset path>
Override the train config to execute multi-GPU training:
torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/multi_gpu_train.json']"
Please note that the distributed training-related options depend on the actual running environment; thus, users may need to remove --standalone , modify --nnodes , or do some other necessary changes according to the machine used. For more details, please refer to pytorch’s official tutorial
.
Override the train config to execute evaluation with the trained model:
python -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json']"
Override the train config and evaluate config to execute multi-GPU evaluation:
torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json','configs/multi_gpu_evaluate.json']"
Execute inference:
python -m monai.bundle run --config_file configs/inference.json
Export checkpoint to TensorRT based models with fp32 or fp16 precision:
python -m monai.bundle trt_export --net_id network_def --filepath models/model_trt.ts --ckpt_file models/model.pt --meta_file configs/metadata.json --config_file configs/inference.json --precision <fp32/fp16> --dynamic_batchsize "[1, 4, 8]" --use_onnx "True" --use_trace "True"
Execute inference with the TensorRT model:
python -m monai.bundle run --config_file "['configs/inference.json', 'configs/inference_trt.json']"
References
[1] Xia, Yingda, et al. “3D Semi-Supervised Learning with Uncertainty-Aware Multi-View Co-Training.” arXiv preprint arXiv:1811.12506 (2018). https://arxiv.org/abs/1811.12506 .
[2] Kerfoot E., Clough J., Oksuz I., Lee J., King A.P., Schnabel J.A. (2019) Left-Ventricle Quantification Using Residual U-Net. In: Pop M. et al. (eds) Statistical Atlases and Computational Models of the Heart. Atrial Segmentation and LV Quantification Challenges. STACOM 2018. Lecture Notes in Computer Science, vol 11395. Springer, Cham. https://doi.org/10.1007/978-3-030-12029-0 _40
License
Copyright (c) MONAI Consortium
Licensed under the Apache License, Version 2.0 (the “License”); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an “AS IS” BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
Follow AI Models on Google News
An easy & free way to support AI Models is to follow our google news feed! More followers will help us reach a wider audience!
Google News: AI Models
