A simultaneous segmentation and classification of nuclei within multitissue histology images based on CoNSeP data
A pre-trained model for simultaneous segmentation and classification of nuclei within multi-tissue histology images based on CoNSeP data. The details of the model can be found in [1].
The model is trained to simultaneously segment and classify nuclei, and a two-stage training approach is utilized:
- Initialize the model with pre-trained weights, and train the decoder only for 50 epochs.
- Finetune all layers for another 50 epochs.
There are two training modes in total. If “original” mode is specified, [270, 270] and [80, 80] are used for patch_size and out_size respectively. If “fast” mode is specified, [256, 256] and [164, 164] are used for patch_size and out_size respectively. The results shown below are based on the “fast” mode.
In this bundle, the first stage is trained with pre-trained weights from some internal data. The original author’s repo and torchvison also provide pre-trained weights but for non-commercial use. Each user is responsible for checking the content of models/datasets and the applicable licenses and determining if suitable for the intended use.
If you want to train the first stage with pre-trained weights, just specify --network_def#pretrained_url <your pretrain weights URL> in the training command below, such as ImageNet
.
Data
The training data is from https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet/ .
- Target: segment instance-level nuclei and classify the nuclei type
- Task: Segmentation and classification
- Modality: RGB images
- Size: 41 image tiles (2009 patches)
The provided labelled data was partitioned, based on the original split, into training (27 tiles) and testing (14 tiles) datasets.
You can download the dataset by using this command:
wget https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet/consep_dataset.zip
unzip consep_dataset.zip
Preprocessing
After download the datasets
, please run scripts/prepare_patches.py to prepare patches from tiles. Prepared patches are saved in <your concep dataset path> /Prepared. The implementation is referring to https://github.com/vqdang/hover_net
. The command is like:
python scripts/prepare_patches.py --root <your concep dataset path>
Training configuration
This model utilized a two-stage approach. The training was performed with the following:
- GPU: At least 24GB of GPU memory.
- Actual Model Input: 256 x 256
- AMP: True
- Optimizer: Adam
- Learning Rate: 1e-4
- Loss: HoVerNetLoss
- Dataset Manager: CacheDataset
Memory Consumption Warning
If you face memory issues with CacheDataset, you can either switch to a regular Dataset class or lower the caching rate cache_rate in the configurations within range [0, 1] to minimize the System RAM requirements.
Input
Input: RGB images
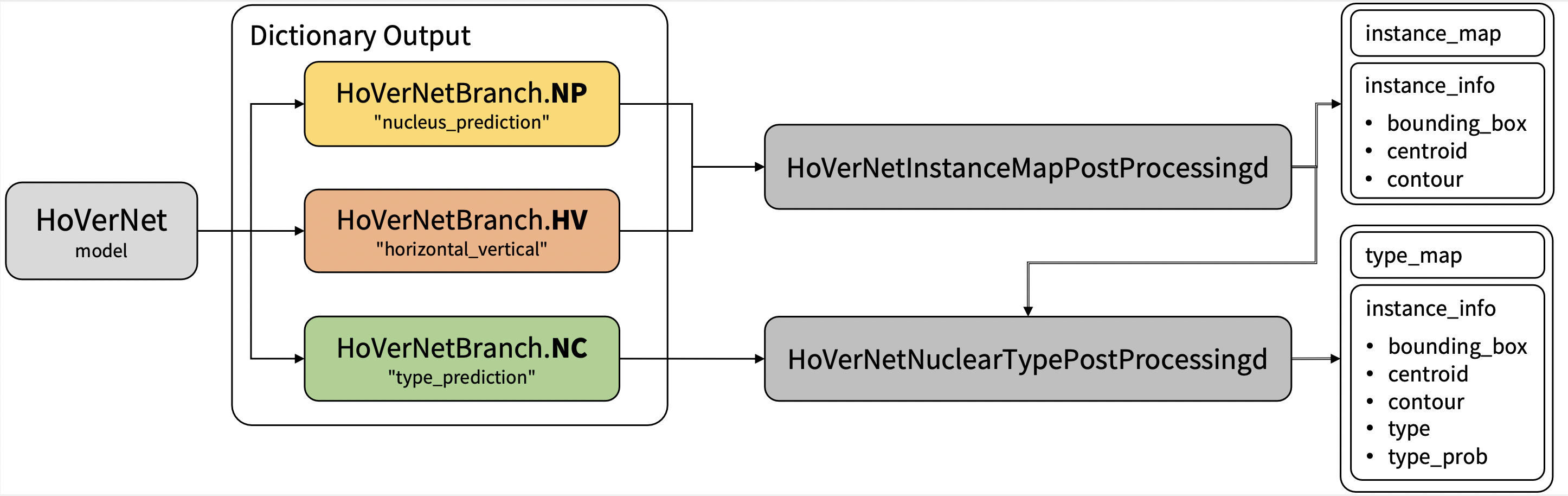
Output
Output: a dictionary with the following keys:
- nucleus_prediction: predict whether or not a pixel belongs to the nuclei or background
- horizontal_vertical: predict the horizontal and vertical distances of nuclear pixels to their centres of mass
- type_prediction: predict the type of nucleus for each pixel
Performance
The achieved metrics on the validation data are:
Fast mode: - Binary Dice: 0.8291 - PQ: 0.4973 - F1d: 0.7417
Note: - Binary Dice is calculated based on the whole input. PQ and F1d were calculated from https://github.com/vqdang/hover _net#inference. - This bundle is non-deterministic because of the bilinear interpolation used in the network. Therefore, reproducing the training process may not get exactly the same performance. Please refer to https://pytorch.org/docs/stable/notes/randomness.html#reproducibility for more details about reproducibility.
Training Loss and Dice
stage1: 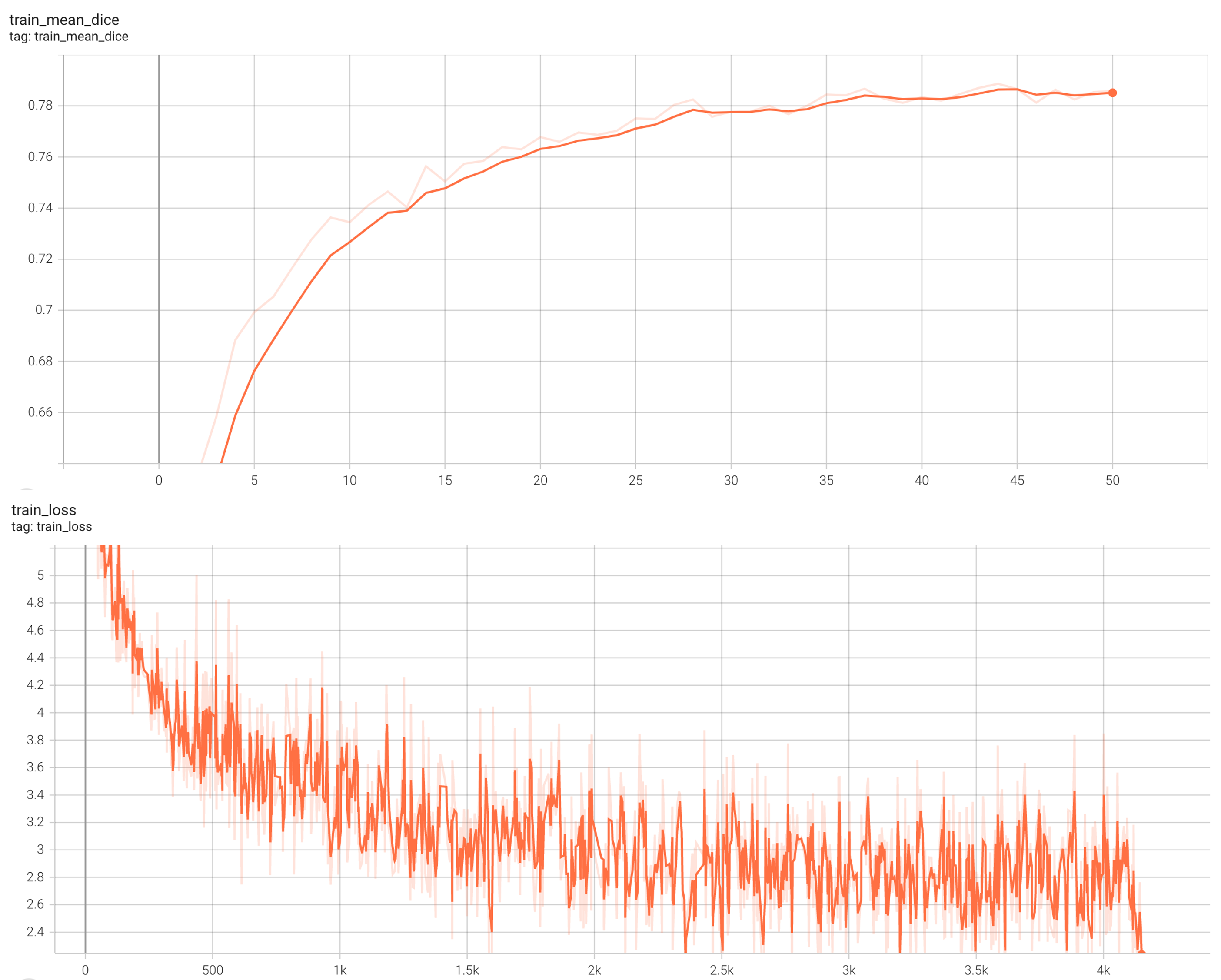
stage2: 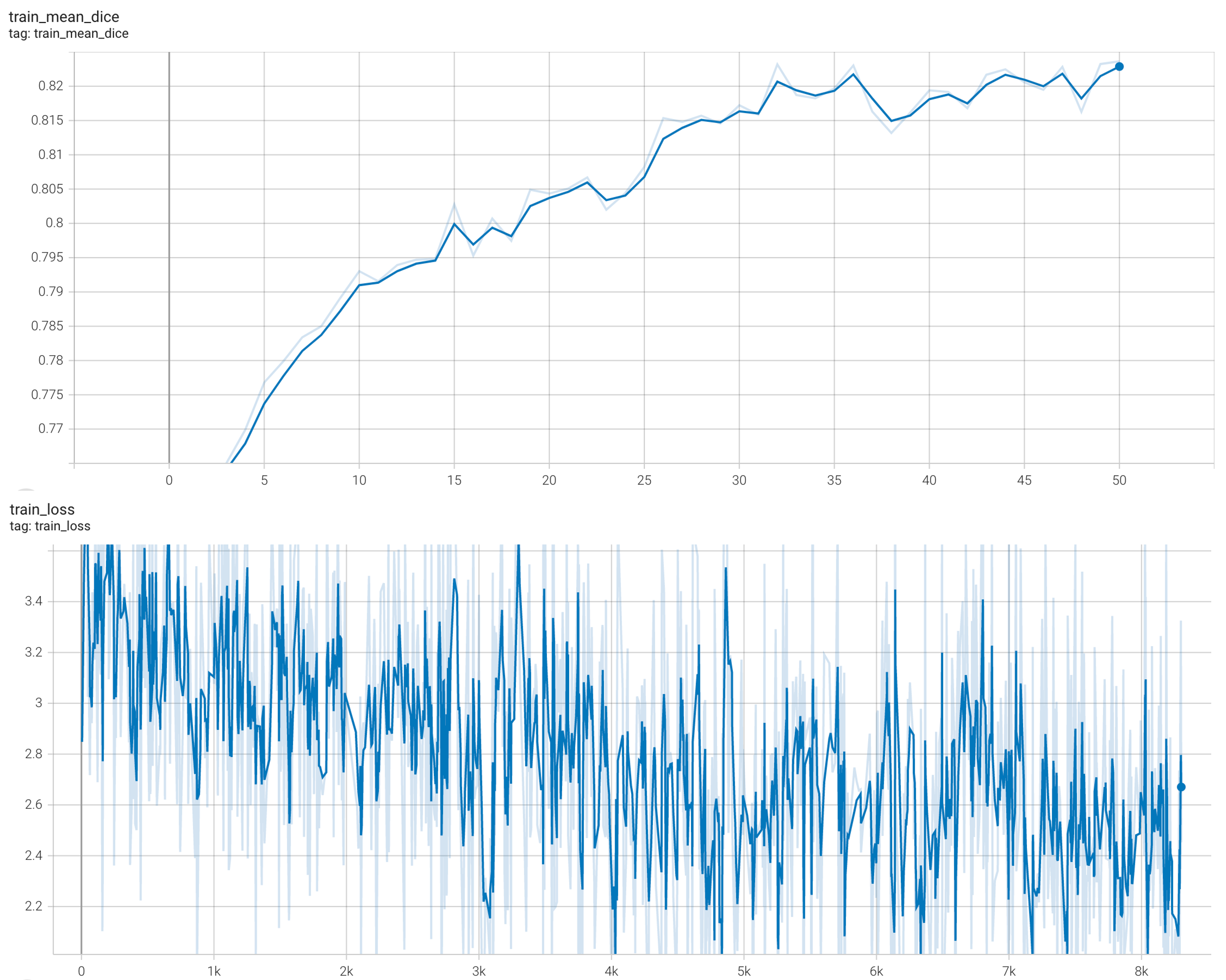
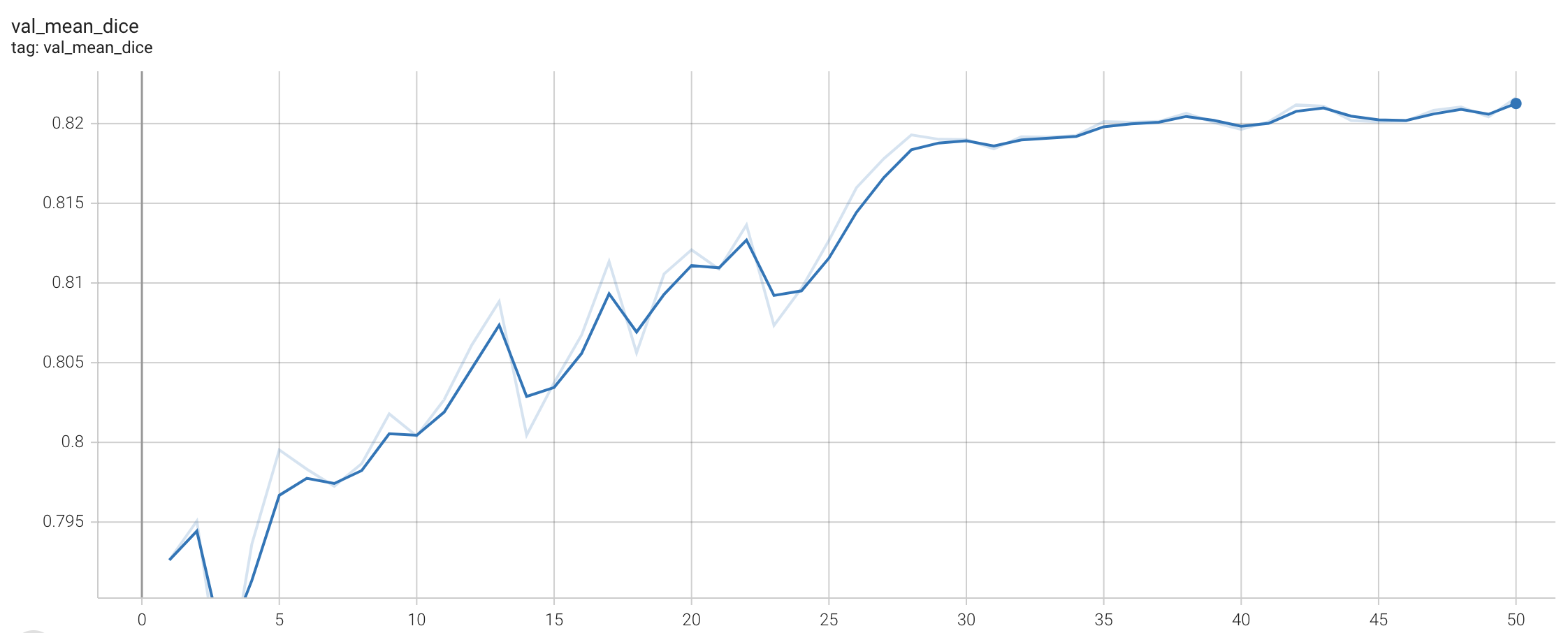
Validation Dice
stage1:
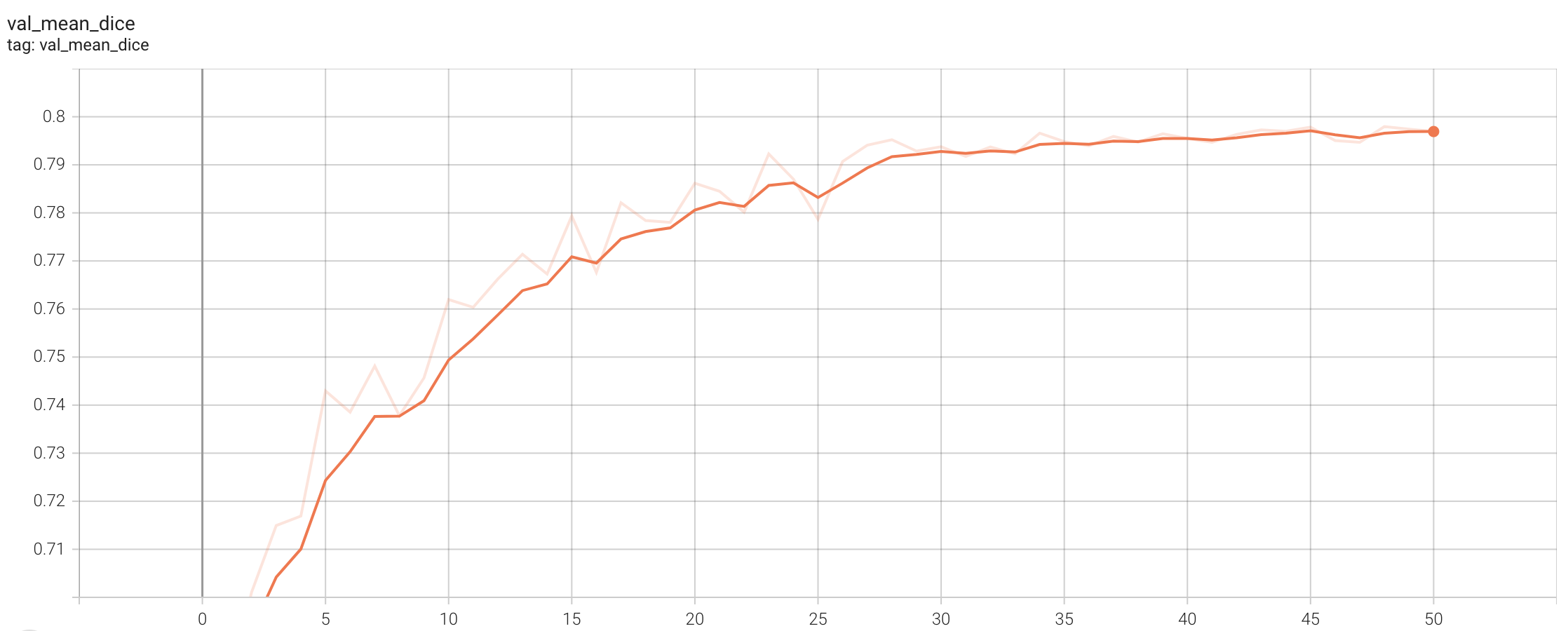
stage2:
MONAI Bundle Commands
In addition to the Pythonic APIs, a few command line interfaces (CLI) are provided to interact with the bundle. The CLI supports flexible use cases, such as overriding configs at runtime and predefining arguments in a file.
For more details usage instructions, visit the MONAI Bundle Configuration Page .
Execute training, the evaluation during the training were evaluated on patches:
Please note that if the default dataset path is not modified with the actual path in the bundle config files, you can also override it by using --dataset_dir :
-
Run first stage
python -m monai.bundle run –config_file configs/train.json –stage 0 –dataset_dir
-
Run second stage
python -m monai.bundle run –config_file configs/train.json –network_def#freeze_encoder False –stage 1 –dataset_dir
Override the train config to execute multi-GPU training:
-
Run first stage
torchrun –standalone –nnodes=1 –nproc_per_node=2 -m monai.bundle run –config_file “[‘configs/train.json’,‘configs/multi_gpu_train.json’]” –batch_size 8 –network_def#freeze_encoder True –stage 0
-
Run second stage
torchrun –standalone –nnodes=1 –nproc_per_node=2 -m monai.bundle run –config_file “[‘configs/train.json’,‘configs/multi_gpu_train.json’]” –batch_size 4 –network_def#freeze_encoder False –stage 1
Override the train config to execute evaluation with the trained model, here we evaluated dice from the whole input instead of the patches:
python -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json']"
Execute inference:
python -m monai.bundle run --config_file configs/inference.json
References
[1] Simon Graham, Quoc Dang Vu, Shan E Ahmed Raza, Ayesha Azam, Yee Wah Tsang, Jin Tae Kwak, Nasir Rajpoot, Hover-Net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images, Medical Image Analysis, 2019 https://doi.org/10.1016/j.media.2019.101563
License
Copyright (c) MONAI Consortium
Licensed under the Apache License, Version 2.0 (the “License”); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an “AS IS” BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
Follow AI Models on Google News
An easy & free way to support AI Models is to follow our google news feed! More followers will help us reach a wider audience!
Google News: AI Models

